Evaluating Cloud Bigtable
(Part 2/4)
In the previous article, we presented the challenges that prompted us to migrate away from HBase on Amazon Web Services (AWS). In this article, we describe how we went about evaluating Cloud Bigtable as a potential datastore for Sift. We did this by assessing Bigtable’s ability to handle the online data load from Sift, by exploring the availability of Google Cloud Platform (GCP) tools to migrate historic data from HBase, and by mimicking HBase operations on Bigtable to surface any potential issues.
The target audience of this article is technology enthusiasts and engineering teams exploring Cloud Bigtable.
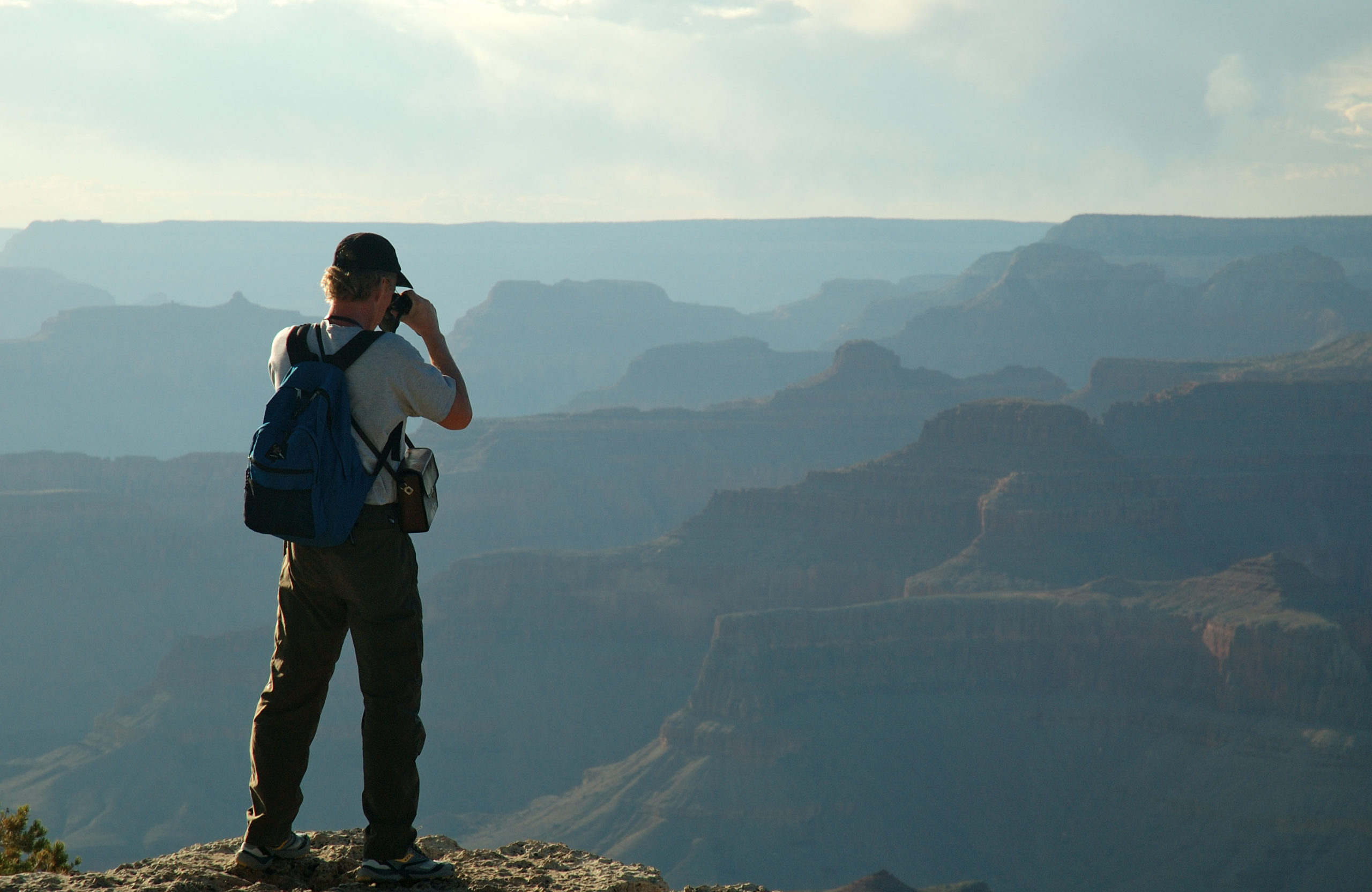

Platform details
By Winter 2018, the Sift platform handled thousands of scoring requests per second from customers around the globe who wanted to prevent fraud on their digital platforms. We were also continuously ingesting data from our customers, which is critical for the correctness of our fraud event scores, at the rate of tens of thousands of requests per second. Both of these translated to about 500,000 requests/s to our HBase datastores, which amounted to a throughput of about 2GB/s for reads and 1GB/s for writes.
For this article, we use a simplified view of our architecture which looked like this in Winter 2018.
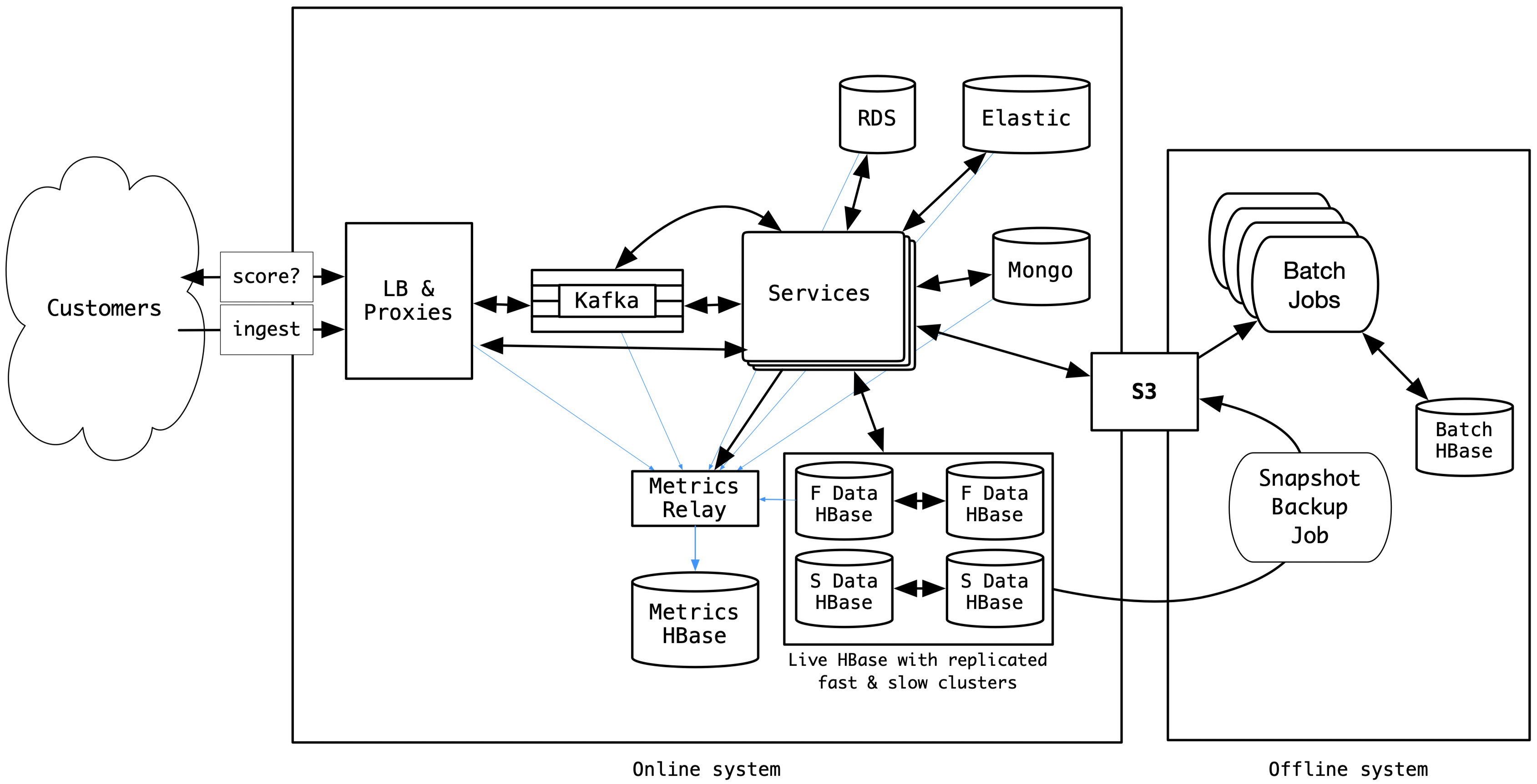
Salient characteristics of our HBase datastore
- There were 100+ tables, with sizes ranging from a few MBs to more than 75TB.
- We had HBase setup in separate slow(HDD) and fast(SSD) tiered clusters; tables that were not used in the online scoring path were kept in the slower and cheaper HDD clusters.
- In each tier, we had two replicated HBase clusters in a hot-standby setup to provide a high availability.
- We had around 1PB of data in our HBase setup that was growing really fast with the business growth.
- The daily snapshots of HBase tables, copied into S3, served as inputs to the business operation batch jobs and disaster recovery.
Operational requirements
- All services must be able to access any table in our main datastore, HBase.
- Our scoring APIs must continue to function, keeping our 99p latency within SLA throughout any operation.
- Our data ingest APIs must be always available, and the internal systems that write the ingested data to the datastore must not miss writing any data.
Proof of concept — Setup
GCP had a lot of useful documentation that helped us understand their cluster management concepts, APIs, administration, cost analysis, and debugging tools. In our initial cost analysis, just from a data storage point of view, Cloud Bigtable seemed significantly cost-effective, financially and operationally, than running HBase ourselves.
That said, before embarking on this migration, we wanted to be sure that “Cloud Bigtable as a service” was robust enough to keep up with Sift’s growing operational load with the growth of the business. We also wanted to have a high confidence with our internal engineering teams that such a migration effort was likely to succeed. With the help of a credit we received from Google, we decided to do a proof of concept (POC) exercise in the Winter of 2018.
POC goals and success criteria
- Ensure that we can read & write to Bigtable at rate & latency that is comparable to what we do with HBase.
- Ensure that we can backfill historic data into Bigtable tables without impacting the performance of our online system.
- Ensure that we can export data from all our tables to a snapshot-like output in reasonable time, for our batch jobs to use.
- Confirm that the cluster replication in Bigtable is fast enough to allow failovers anytime.
- Evaluate Bigtable scaling, and verify that we could scale Bigtable without violating our SLAs.
Datacenter selection
Our application services and the EC2 instances hosting HBase/HDFS were running on AWS us-east-1 region. Since we wanted to minimize the latency effects during migration, we picked the geographically nearby GCP region, the us-east4.
Mirroring module
In order to load-test Bigtable, we decided to mirror our online traffic to Bigtable. This required us to change our code in just one module. All our application services and batch jobs accessed HBase via a Java DB connector object that maintained connections to our HDD and SSD tiers. We modified this object to make additional connections, using the Cloud Bigtable HBase client for Java, to the corresponding Bigtable tier in our GCP POC project.
Almost all our table accesses in Java were done using the HTableInterface for a table. Therefore, we created a new implementation of HTableInterface (say, BigtableMirror) that wrapped the actual HBase table’s HTableInterface as a delegate. The BigtableMirror also contained the connection to Bigtable. The overridden methods in BigtableMirror would mirror a percentage of the HBase traffic to Bigtable, and collect metrics around successful and failed mirror writes and mirror reads. We used a Zookeeper node to hold information about which service to enable mirroring, and what percentage of traffic to mirror to Bigtable. In our application services, we used the Apache Curator client to listen to changes to this Zookeeper node, so we could modify traffic mirroring percentages easily by adjusting the Zookeeper node value.
Backfill pipeline
Copying historic data from HBase to Bigtable was possible with the open source Google Dataflow code that can be set to import Hadoop sequence files to Bigtable table. The sequence files needed to be present in Google Cloud Storage (GCS). In order to achieve this, we wrote a collection of tools that can be run in a pipeline, as shown below, to backfill historic data from HBase (on AWS) to Bigtable.
We built a collection of tools to,
- create Bigtable tables with splits for non-ascii rowkeys.
- create Bigtable table with schema matching an existing HBase table; this does the conversion from versions or TTL (in HBase table schema) to GC policy (in Bigtable table schema) when creating matching columns families in Bigtable.
- convert hFiles (from S3 snapshots) to Hadoop sequence-files. GCP documents suggested to use org.apache.hadoop.hbase.mapreduce.Export from a namenode, but we didn’t want the reads to put pressure on the live system, so we modified the Export to read from S3 snapshot.
- automate data transfer from this new S3 bucket into a GCS bucket.
- import GCS sequence-files into a given Bigtable table.
- export a Bigtable table to GCS sequence files, for our batch jobs.
We also built a pipeline orchestrator that used the above tools in different combinations with different parameters (eg: how many tables to process in parallel, configuring per-table number of export mappers, memory & disk per mapper, number of dataflow workers, memory & disk for worker, number of table splits, etc.).
Synthetic load-test
We also ran a synthetic load-test (from google-cloud-go) for a few weeks to make sure the Bigtable clusters can sustain high traffic with different load patterns for an extended period of time.
With this framework, we studied latency, throughput, system stability, and capacity requirement characteristics for mirroring, exports and backfills over a 2 month period.
Proof of concept — Findings
Some of the interesting findings from our POC are listed below. Keep in mind that some of these may have been addressed by Google in newer releases or documentation updates.

Better Bigtable tools
The first thing we liked in Bigtable was the ease with which we could failover between replica clusters. This had been an operational nightmare for us in HBase world.
In HBase, we did not have clear visibility into read/write rates of tables. With BigTable, we could identify tables with high request rate, tables with high throughput, per-table latencies, per-table errors and so on very easily, as shown below:
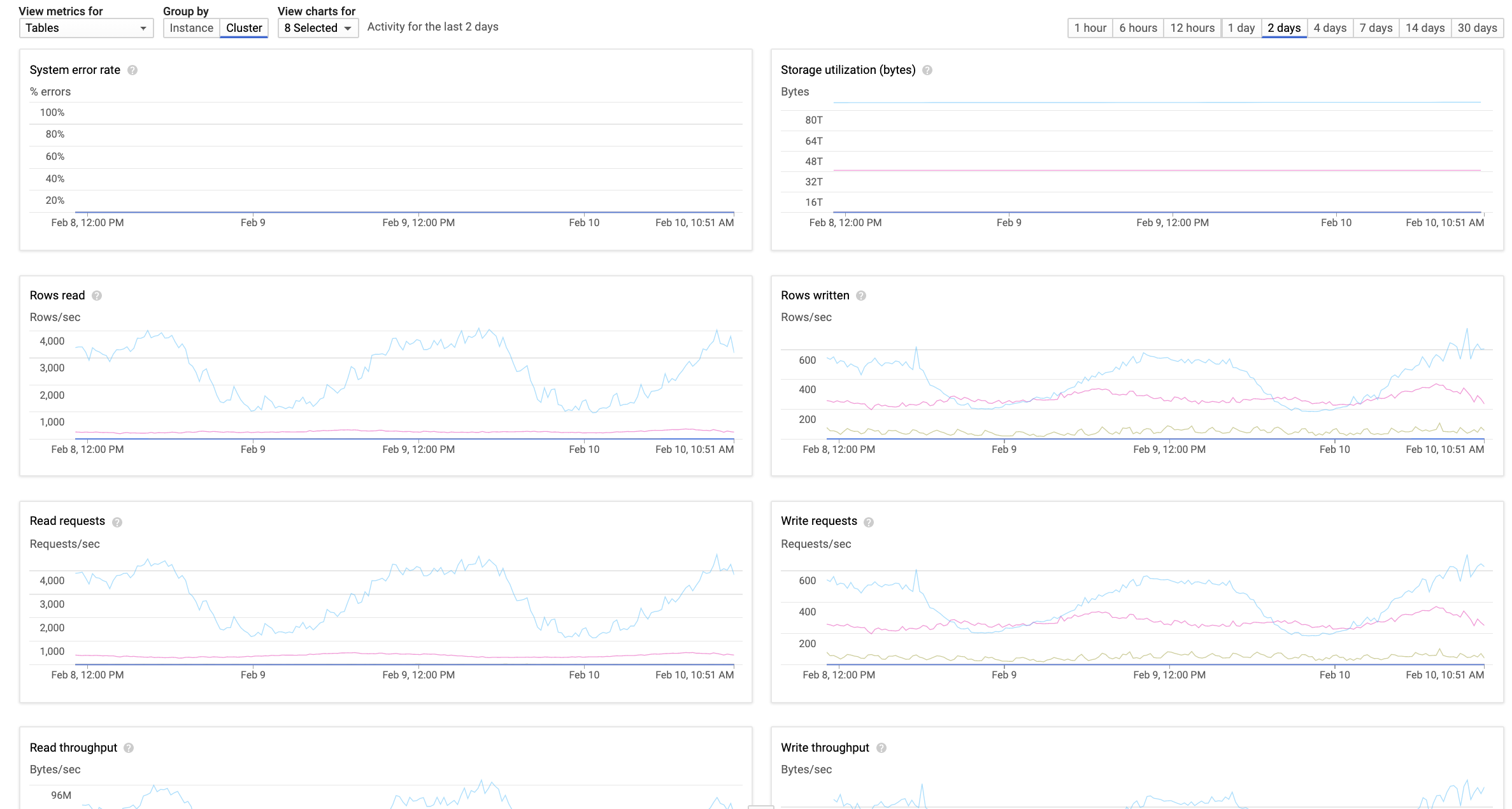
Cloud Bigtable also provided numerous Bigtable insights via Stackdriver metrics and a fantastic tool called Key Visualizer that gave us a lot of visibility into data distribution, hotkeys, large rows etc. for each table.
It was also fairly easy to setup a Dataflow job to import or export Bigtable data.
Documentation shortcomings
In the hFile to sequencefile export job, we wanted to enable compression to minimize storage & transfer cost. We initially tried with bzip2 and Snappy, but only a sequencefile with GZIP compression was recognized by the Dataflow import code. This was not clearly documented.
We pre-split tables while creating them in HBase in order to parallelize and improve performance. Our rowkeys are also not ASCII. We could not get the “cbt createtable ..” command to create table with splits, with non-ascii rowkeys, due to lack of documentation and support. To workaround, we had to create tables using a Python script that used the google-cloud-bigtable and binascii python modules to specify non-ascii split keys.
API/Implementation differences
Google has documented API incompatibilities between Java hbase client and Cloud Bigtable HBase client for Java. But there were other subtle differences that we saw. For instance, in the “Result get(Get get)” and “Result[] get(List<Get> gets)” APIs, when there were no results, the hbase client returned an empty Result object. But the Cloud Bigtable HBase client returned null.
Bigtable scans were a bigger problem for us. In hbase-client on HBase, we could do setBatching/setCaching to control the prefetching behavior of the client. Bigtable hbase client seemed to ignore these settings and was doing its own pre-fetching. This was an issue for some of our tables where the scans unintentionally became unbounded, resulting in too much data being read from Bigtable. In addition to putting excessive load on the Bigtable cluster, this also caused a lot of unnecessary and costly data transfer across the clouds in our POC. For the POC we excluded such tables, but allocated time for fixing the scan issue as part of the migration.
Performance tweaks
We wanted to be able to import sequence files to the Bigtable tables as aggressively as possible without affecting client & Bigtable system performance. We also wanted to be able to export the table data as sequence files for all tables within 24 hours because this would be the input to the daily batch jobs.
As per the GCP documentation, it was reasonable to run the import Dataflow job with workers = 3xN where N is the number of Bigtable nodes in a cluster. However we were barely able to run import jobs with 1xN workers; anything more than that was causing cluster CPU utilization to be very high. For the export Dataflow job, we were able to push it to about 7xN workers (as opposed to the recommended 10xN) without causing high CPU utilization.
In HBase we used replication; we wanted to maintain the same high-availability pattern in Bigtable. Therefore our POC tests were run on a replicated Bigtable instance, with single-cluster routing in order to be highly consistent. To control the high CPU utilization during aggressive imports/exports, we decided to do the import/export from the cluster that is not serving live traffic. We created different application profiles with different single-cluster routing to achieve this. But even with this setup, there was often an increase in hot node CPU in the live cluster when importing aggressively in the secondary replica.
Later, Google engineers advised us to set a mutationThresholdMs parameter in the Dataflow import job to make imports reasonably well behaved, but this slowed down the time it took to import.
Large rows
Bigtable imposes hard limits on cell size and row size. HBase didn’t have this constraint. Since we were liberal in our TTLs and versions in HBase, we feared that our data might not comply with Bigtable limits. To check this, we added counters in our hFiles to sequence files Export job. The counters revealed that we had a fraction of our tables, some of them huge, with rows and cells exceeding this limit. Since we caught this issue in the POC, we could allocate time budget to fix it in our migration planning.
Other quirks
Even though it was easy to setup a Dataflow import/export job for Bigtable, the lack of a progress indicator made it hard to plan. Some of the large tables took more than 2 days to import, and we didn’t know how long to wait.
In HBase land, we used the built-in snapshot feature of HBase to take a snapshot, and backup the snapshot data to S3 in an incremental manner. This gave us a lot of space savings (because day 2 data only had the new edits on top of day 1 data). But with Bigtable, the only way to export the table data as a snapshot was by reading out all the table data as sequence files. This was quite expensive because every day we ended up creating a new dataset of all tables.
We had picked the us-east4 regions in GCP due to its proximity with the AWS region we were on to minimize inter-cloud latency during migration; but this GCP region was resource limited in Winter 2018. As a result, we could not balloon the Bigtable cluster to hundreds of nodes to run the import, and then scale it down. Since the POC credit from Google was running out, we had to import the data into a large Bigtable cluster in us-east1 region, and then do an inter-region replication to bring the data over to us-east4. The inter-region replication, though, was lightning fast.
The mirroring experiment was successful, but we had to exclude three high throughput tables from our POC test because they were polluting the results (when we did the actual migration, we handled these tables differently).
Post evaluation thoughts
In this article, we described how we did the evaluation work for our migration. Our approach was to collect as many data points as possible, using the GCP trial credit. During the POC period, the GCP program managers were extremely helpful to clarify issues and give suggestions.
We were satisfied during the POC that we were able to mirror 100% of our HBase traffic to Bigtable without experiencing any back-pressure issues in our application services. Our other success criteria around load, backfill, backup, failover, and scaling were also met.
Overall, the POC was very promising. We liked the extra visibility we got into the performance and utilization of Bigtable and the stability of the system. The findings helped us to convince our organization that a zero-downtime migration is feasible and beneficial to our business.
What is next?
- Planning the zero-downtime data migration: let’s ensure there is zero customer impact during the migration.
- Lessons learned from data migration: key takeaways and lessons learned during this journey.

